Medical doctors who specialize in rare diseases get only so many opportunities to learn as they go. The lack of diverse healthcare data to train students is a key challenge in these fields.
“When you are working in a setting with scarce data, your performance correlates with experience — the more images you see, the better you become,” said Christian Bluethgen, a thoracic radiologist and Stanford Center for AI in Medicine and Imaging (AIMI) postdoc researcher who has studied rare lung diseases for the last seven years.
When Stability AI released Stable Diffusion, its text-to-image foundation model, to the public in August, Bluethgen had an idea: What if you could combine a real need in medicine with the ease of creating beautiful images from simple text prompts? If Stable Diffusion could create medical images that accurately depict the clinical context, it could alleviate the gap in training data.
Bluethgen teamed up with Pierre Chambon, a Stanford graduate student at the Institute for Computational and Mathematical Engineering and machine learning (ML) researcher at AIMI, to design a study that would seek to expand the capabilities of Stable Diffusion to generate the most common type of medical images — chest X-rays.
Together, they found that with some additional training, the general-purpose latent diffusion model performed surprisingly well at the task of creating images of human lungs with recognizable abnormalities. It’s a promising breakthrough that could lead to more widespread research, a better understanding of rare diseases, and possibly even development of new treatment protocols.
From general-purpose to domain-specific
Until now, foundation models trained in natural images and language have not performed well when given domain-specific tasks. Professional fields such as medicine and finance have their own jargon, terminology, and rules, which are not accounted for in general training datasets. But one advantage presented itself for the team’s study: Radiologists always prepare a detailed text report that describes their findings in each image they analyze. By adding this training data into their Stable Diffusion model, the team hoped that the model could learn to create synthetic medical imaging data when prompted with relevant medical keywords.
“We are not the first to train a model for chest X-rays, but previously you had to do it with dedicated datasets and pay a very high price for the compute power,” said Chambon. “Those barriers prevent a lot of important research. We wanted to see if you could bootstrap the approach and use the existing open-source foundation model with only minor tweaks.”
Three-step process
To test Stable Diffusion’s capabilities, Bluethgen and Chambon examined three sub-components of the model’s architecture:
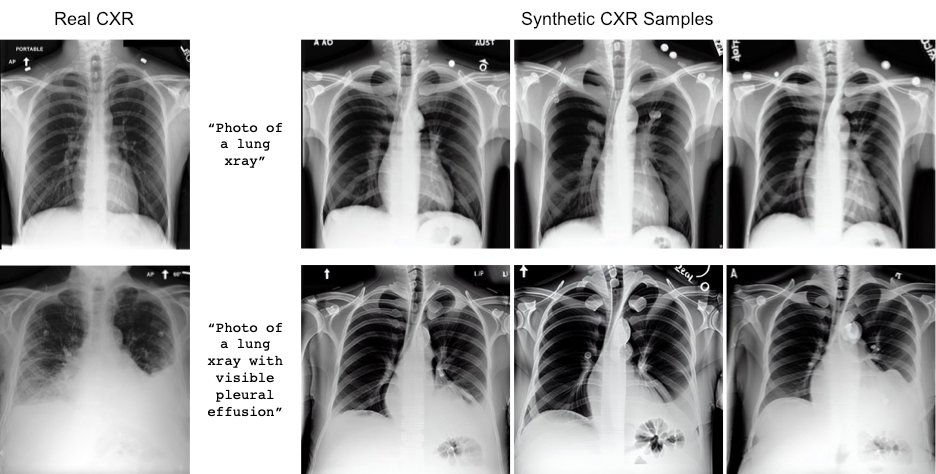
The researchers created a dataset to study the image autoencoder and text encoder components. They randomly selected 1,000 frontal radiographs from each of two large, public datasets, called CheXpert and MIMIC-CXR. Then they added five hand-selected images of normal chest X-rays and five images featuring a clearly visible abnormality (in this case, fluid build-up between tissues, called a pleural effusion).
These images were paired with a set of simple text prompts for testing various ways of fine-tuning the components. Finally, they pulled a sample of 1 million general text prompts from the LAION-400M open dataset, (a large-scale, non-curated set of image-text pairs designed for model training and broad research purposes).
Key findings
Here is what they asked and found, at a high level:
Text Encoder: Using CLIP, a general domain neural network from Open AI that connects text and images, could the model generate a meaningful result when given a text prompt like “pleural effusion” that is specific to the field of radiology? The answer was yes — the text encoder on its own provided sufficient context for the U-Net to create medically accurate images.
VAE: Could the Stable Diffusion autoencoder trained on natural images successfully present a medical image after it had been un-compressed? The result, again, was yes. “Some of the annotations in the original images got scrambled,” said Bluethgen, “so it wasn’t perfect, but taking a first-principles approach, we decided to flag that as an opportunity for a future exploration.”
U-Net: Given the out-of-the-box capabilities of the other two components, could the U-Net create images that are anatomically correct and represent the correct set of abnormalities, depending on the prompt? In this case, Bluethgen and Chambon concluded that additional fine-tuning was needed. “On the first attempt, the original U-Net didn’t know how to generate medical images,” Chambon reports. “But with some additional training, we were able to get to something usable.”
A glimpse of what’s ahead
After experimenting with prompts and benchmarking their efforts using both quantitative quality metrics and qualitative radiologist-driven evaluations, the scholars found their best-performing model could be conditioned to insert a realistic-looking abnormality on a synthetic radiology image while maintaining a 95% accuracy on a deep learning model trained to classify images based on abnormalities.
In follow-up work, Chambon and Bluethgen scaled up training efforts, using tens of thousands of chest X-rays and corresponding reports. The resulting model (called RoentGen, a portmanteau of Roentgen and Generator), announced on Nov. 23, can create CXR images with higher fidelity and increased diversity, and grants a more fine-grained control over image features like size and laterality of the findings through natural language text prompts. (The preprint is available here.)
While this work builds on previous studies, it is the first of its kind to look at latent diffusion models for thoracic imaging, as well as the first to explore the new Stable Diffusion model for generating medical images. Admittedly, several limitations surfaced as the team reflected on the approach:
Additionally, even if this model someday worked perfectly, it’s unclear if medical researchers could legally use it. Stable Diffusion’s open-source license agreement currently prevents users from generating images for medical advice or medical results interpretation.
Art or annotated x-ray?
Despite current limitations, Bluethgen and Chambon say they were amazed at the kind of images they were able to generate from this first phase of research.
“Typing a text prompt and getting back whatever you wrote down in the form of a high-quality image is an incredible invention — for any context,” said Bluethgen. “It was mind-blowing to see how well the lung X-ray images got reconstructed. They were realistic, not cartoonish.”
Moving forward, the team plans to explore how powerful latent-diffusion models can learn a wider range of abnormalities, start to combine more than one abnormality in a single image, and eventually extend the research to other kinds of imaging besides X-rays and different body parts.
“There’s a lot of potential in this line of work,” Chambon concludes. “With better medical datasets, we may be able to understand modern disease and treat patients in optimal ways.”
“Adapting Pretrained Vision-Language Foundational Models to Medical Imaging Domains Background” was published in preprint server ArXiv in October. In addition to Bluethgen and Chambon, Curt Langlotz, professor of radiology and faculty affiliate of HAI, and Akshay Chaudhari, assistant professor (research) of radiology, advised and co-authored the study.
Nikki Goth Itoi is a contributing writer for the Stanford Institute for Human-Centered AI.
Welcome to the VentureBeat community!
Our guest posting program is where technical experts share insights and provide neutral, non-vested deep dives on AI, data infrastructure, cybersecurity and other cutting-edge technologies shaping the future of enterprise.
Read more from our guest post program — and check out our guidelines if you’re interested in contributing an article of your own!
